Research
Chemical Origins of Life
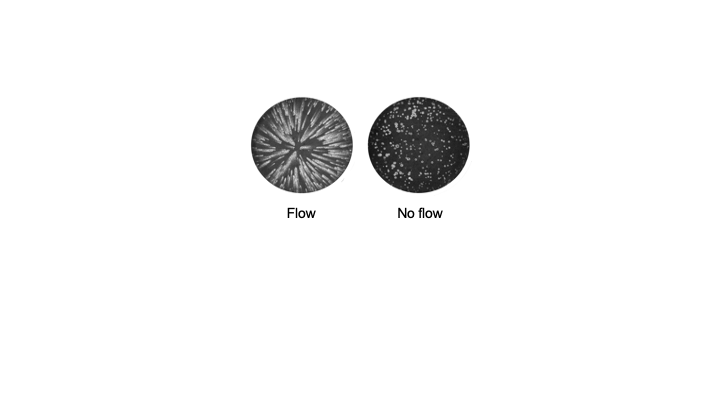
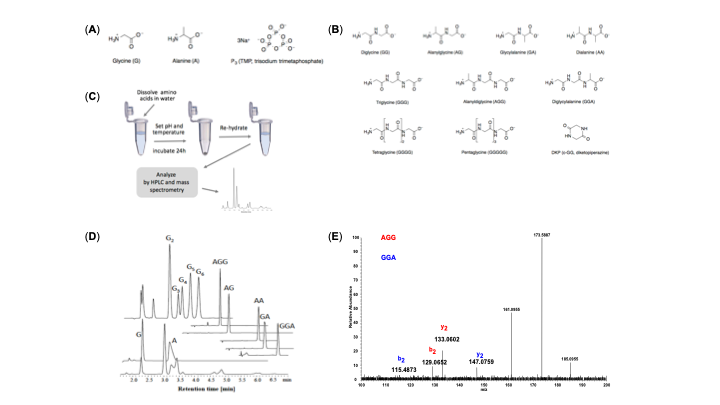
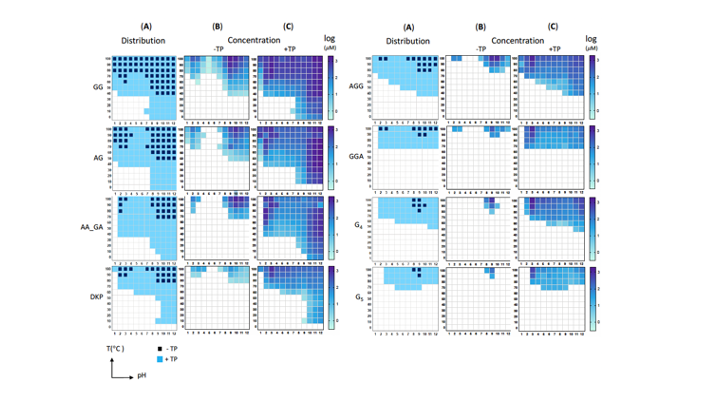
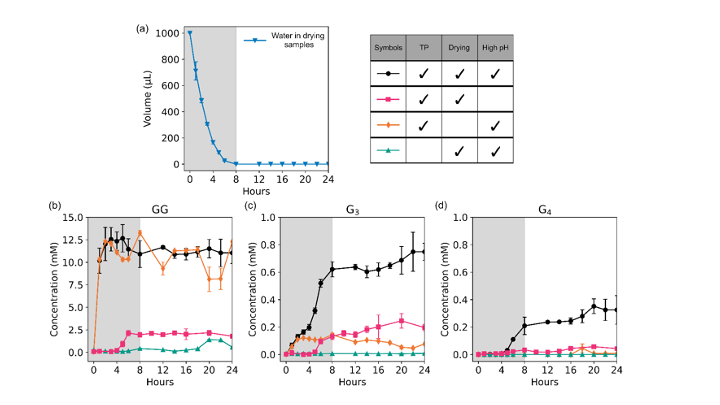
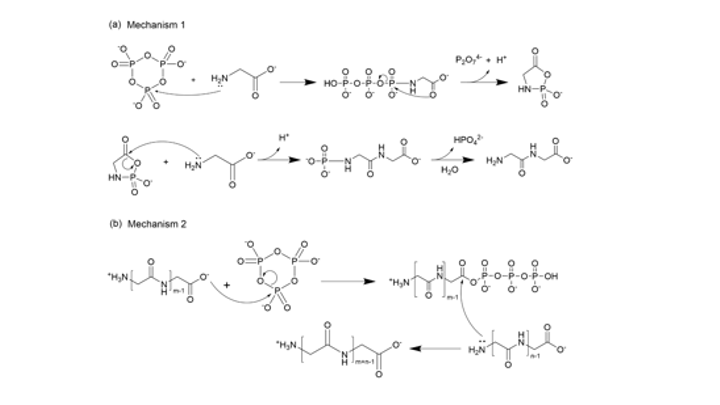
Since the 1953 experiment by Stanley Miller and Harold Urey, we have known that a mixture of simple gases of a primitive early-Earth atmosphere (water vapor, methane, ammonia, hydrogen) could be catalyzed to produce amino acids, the building blocks of life. But possible further steps, where amino acids join to form peptides or proteins with primitive functions are unknown. We have shown how different environmental conditions of pH, heat, and drying can promote the formation of different strings of amino acids (peptides), a step toward linking synthesis environment to the emergence of molecular information (Sibilska, 2018). Others have suggested how short peptides could interact to serve as information or replication templates, related to amyloid formation in neurodegenerative process that are linked to Alzheimers or Huntington’s disease. Meanwhile, clever biochemists have figured out ways to engineer peptides that can perform feats of self-replication. But it remains a wide-open challenge how a de novo wet-lab system, starting with amino acids, simple activating agents, and interfaces (solid-liquid or liquid-liquid) surfaces can give rise to any form of self-replication.
The Human Virome in Health and Disease
The term, microbiome, describes the collective microbes that inhibit our bodies and other environments. In the Yin Lab we focus on a subset of the microbiome, the virome, or collective viruses associated with humans and their environments. Some viruses might be beneficial; for example, viruses that infect pathogenic microbes may contribute to a microbiome that promotes our health. Other viruses, especially the ones that make us sick, are well known, for example, HIV-1, hepatitis, ebola, the common cold, zika, and influenza. More recently, the COVID-19 pandemic was caused by the severe acute respiratory syndrome coronavirus 2, known as SARS-CoV-2.
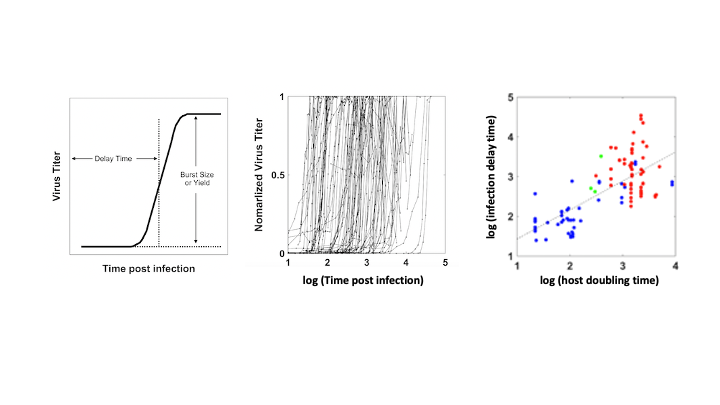
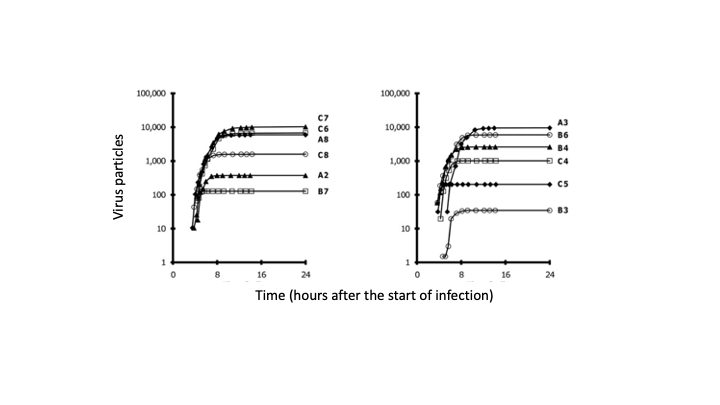
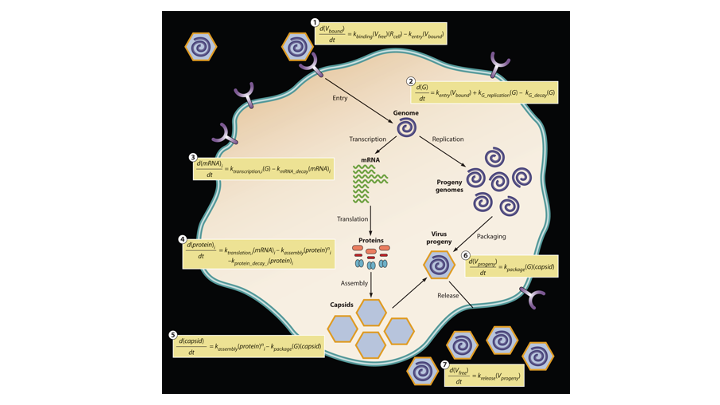
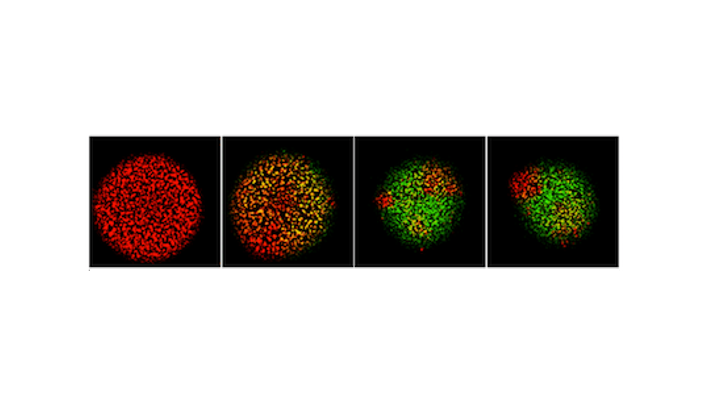
Many viruses, including coronaviruses, make errors when they reproduce by introducing mutations in their genomes. Such mutations can expand their powers, enabling them to infect new host species, like jumping from bats to humans. Or the mutant viruses can evade host immune responses and escape drug treatments. More broadly, the mutations can lead to diseases that persist or re-emerge.
Alternatively, extreme errors can delete key virus functions, rendering the virus non-infectious. Such ‘dead’ viruses are ubiquitous byproducts of virus infections in humans and in natural host reservoirs. Remarkably, such dead viruses can spring to life if they invade a productively infected host cell, where they can reproduce at the expense of the normal virus. These molecular parasites of normal virus growth have potential as therapeutics. It is currently unknown to what extent such defective viruses might stimulate host immune responses in ways that might be protective versus causing more severe disease. In the Yin Lab we are working to understand how such defective viruses can impact disease severity and how they might be engineered to prevent future pandemics.
Human Physiology
In the healthy human, diverse organ systems take up and distribute oxygen, digest food for energy, and rid the body of waste. Such systems are coordinated in part by the nervous system, which senses, transmits, and reacts to chemical and electrical signals throughout the body. But as individuals age, their physiological functions can decline. For example, 60% of adults over 40 suffer symptoms of the lower urinary tract, which include a need to urinate that may be frequent, urgent, or at night. Methods of nerve stimulation have potential to treat such symptoms, but nerve signaling from and to the bladder and urethra, while measurable, are not yet understood well enough to guide treatment.
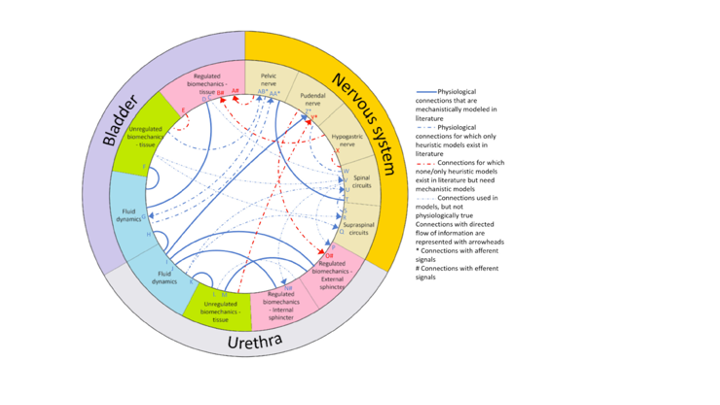
In the Yin Lab we are part of a multi-disciplinary team to advance hybrid mechanistic-and-neural-network computational models of lower urinary tract function guided by wet-lab experiments on rats. As chemical engineers, we are applying principles of dimensional analysis to enable scale-up of results from rats to humans. At the same time, we are pioneering hybrid models that combine the benefits of mechanistic understanding with emerging data-driven methods.
In our cross-disciplinary study of the lower urinary tract, we reviewed the literature, highlighting how fluid flows and bio-mechanical signals share information with the nervous system. Our current efforts are advancing hybrid mechanistic-neural network models, informed by experimental rat data. The greatest gap in our understanding in the interface of the bladder-and-urethra with the nervous